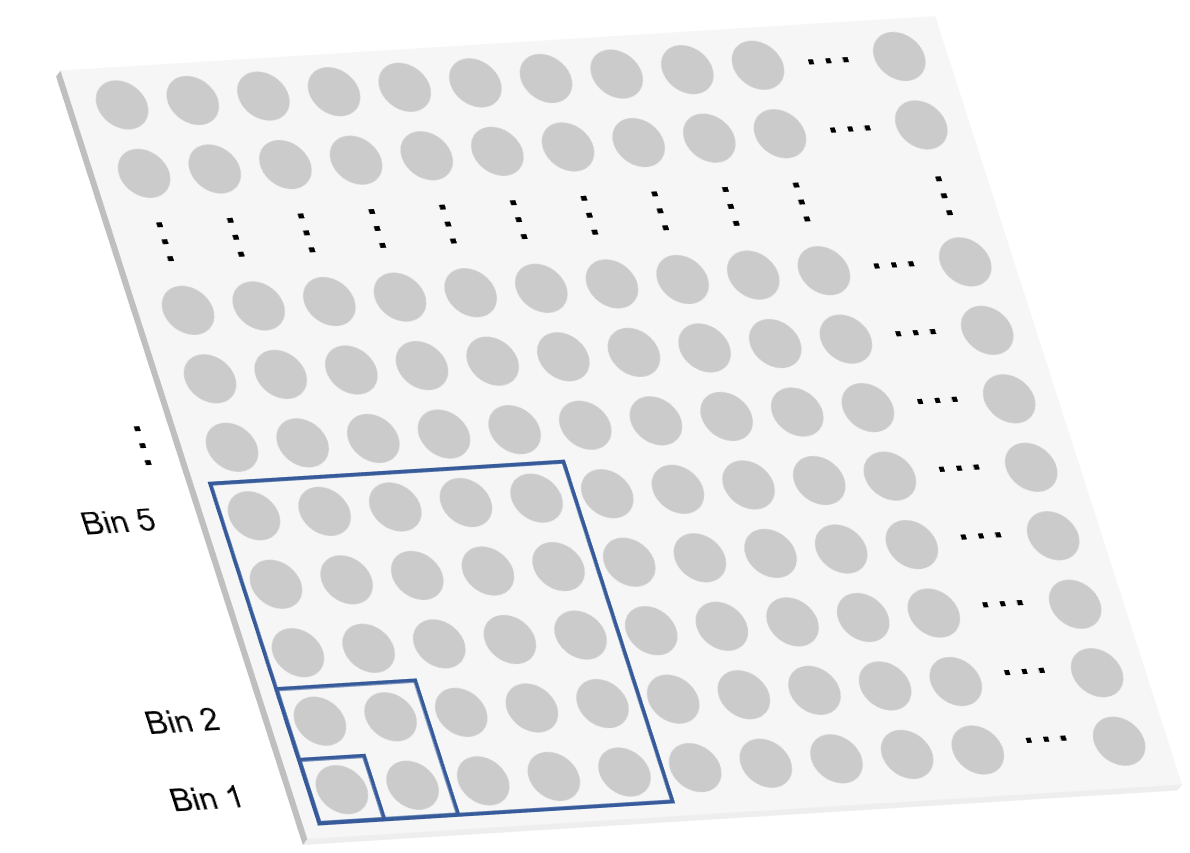
.gefThe feature expression matrix file in HDF5 format for visualization. It contains the MID count for each gene of each spot. A spot is a binning unit that has a fixed-sized square shape in which the expression value in this square is accumulated. By default, a visualization .gef includes spot sizes of bin 1, 5, 10, 20, 50, 100, 150, 200.
.cellbin.gefThe cellbin feature expression matrix file in HDF5 format. It contains the spatial location and area of each cell, the MID count for each gene of each cell, and the cluster the cell belongs to. In .cellbin.gef, the cell is the smallest data unit.
Only available when the cell segmentation was done based on an microscopy image.